gene_token_id list | gene_expression list | cell_barcode string | sample string | num_features int64 | guide_target string | gene_target string | n_genes_by_counts int64 | total_counts float64 | total_counts_mt float64 | pct_counts_mt float64 | pass_guide_filter int64 |
|---|---|---|---|---|---|---|---|---|---|---|---|
[
29,
30,
34,
35,
38,
49,
51,
55,
59,
60,
64,
66,
67,
69,
75,
81,
82,
86,
88,
91,
93,
94,
105,
109,
112,
113,
117,
145,
152,
180,
187,
204,
212,
218,
221,
226,
227,
230,
240,
249,
250,
256,
258,
259,
260,
269,
274,
276,
27... | [
1,
3,
1,
3,
1,
1,
1,
2,
1,
3,
1,
6,
1,
5,
2,
2,
1,
1,
1,
2,
1,
1,
3,
1,
1,
1,
1,
1,
1,
25,
2,
1,
1,
7,
4,
1,
1,
34,
1,
1,
1,
1,
1,
1,
1,
1,
2,
3,
4,
1,
1,
1,
2,
1,
1,
1,
1,
3,
1,
1,
2,
1,
4,
... | AAACCAAAGACATGTT-HCT116_Batch1 | HCT116_Batch1 | 2 | ST14_P1P2-1|ST14_P1P2-2 | ST14 | 4,883 | 19,136 | 1,179 | 6.161162 | 1 |
[
30,
32,
35,
36,
51,
52,
55,
60,
62,
64,
66,
67,
69,
75,
77,
81,
86,
88,
91,
92,
93,
94,
97,
105,
109,
112,
113,
143,
146,
152,
154,
155,
180,
186,
187,
189,
190,
193,
194,
196,
200,
204,
212,
218,
221,
222,
227,
230,
237... | [
9,
1,
5,
11,
10,
2,
1,
3,
5,
1,
12,
5,
7,
1,
2,
4,
4,
3,
2,
1,
5,
11,
1,
3,
2,
7,
1,
2,
2,
5,
1,
2,
41,
1,
6,
1,
3,
1,
2,
2,
3,
5,
1,
20,
18,
3,
13,
94,
2,
1,
1,
2,
4,
3,
6,
1,
1,
8,
2,
10,
7,
1... | AAACCAAAGACCCAAC-HCT116_Batch1 | HCT116_Batch1 | 2 | SIGLEC5_P1P2-1|SIGLEC5_P1P2-2 | SIGLEC5 | 8,130 | 47,916 | 1,562 | 3.259871 | 1 |
[
18,
30,
35,
36,
40,
49,
51,
52,
59,
60,
62,
64,
66,
68,
69,
75,
79,
81,
84,
86,
88,
91,
93,
94,
97,
102,
105,
112,
136,
152,
154,
156,
177,
180,
183,
187,
193,
194,
197,
200,
204,
212,
218,
221,
227,
230,
236,
241,
245,
... | [
1,
5,
3,
3,
1,
2,
3,
1,
1,
4,
3,
2,
9,
1,
7,
2,
3,
5,
1,
2,
3,
2,
2,
4,
2,
2,
2,
1,
2,
2,
1,
1,
1,
33,
2,
3,
2,
1,
1,
3,
3,
2,
19,
7,
2,
53,
1,
1,
2,
2,
1,
1,
1,
1,
3,
2,
6,
2,
3,
4,
1,
4,
7,
... | AAACCAAAGAGGTACG-HCT116_Batch1 | HCT116_Batch1 | 2 | VSNL1_P1P2-1|VSNL1_P1P2-2 | VSNL1 | 6,531 | 28,435 | 1,042 | 3.664498 | 1 |
[12,30,35,36,38,43,49,51,55,60,62,66,67,68,69,75,77,79,81,86,91,94,97,100,105,112,145,146,154,156,18(...TRUNCATED) | [1.0,3.0,4.0,1.0,2.0,1.0,3.0,5.0,2.0,2.0,1.0,10.0,3.0,2.0,6.0,1.0,1.0,2.0,4.0,4.0,1.0,2.0,1.0,1.0,2.(...TRUNCATED) | AAACCAAAGCGATTAT-HCT116_Batch1 | HCT116_Batch1 | 2 | KCNK7_P1P2-1|KCNK7_P1P2-2 | KCNK7 | 5,931 | 26,080 | 1,087 | 4.167945 | 1 |
[30,31,35,36,38,49,51,55,59,60,66,67,68,69,75,77,79,81,82,86,88,93,94,97,100,105,112,136,141,145,146(...TRUNCATED) | [7.0,1.0,2.0,5.0,2.0,1.0,5.0,1.0,3.0,2.0,21.0,1.0,1.0,18.0,1.0,1.0,4.0,4.0,1.0,3.0,3.0,1.0,6.0,1.0,1(...TRUNCATED) | AAACCAAAGGCTTAAT-HCT116_Batch1 | HCT116_Batch1 | 2 | APOA4_P1P2-1|APOA4_P1P2-2 | APOA4 | 7,157 | 38,366 | 955 | 2.489183 | 1 |
[18,30,36,38,49,51,59,60,66,69,81,86,88,91,93,105,112,147,152,154,180,183,192,193,200,204,218,221,22(...TRUNCATED) | [2.0,3.0,2.0,1.0,1.0,1.0,1.0,1.0,2.0,3.0,1.0,1.0,1.0,2.0,1.0,1.0,2.0,1.0,1.0,2.0,9.0,1.0,1.0,1.0,1.0(...TRUNCATED) | AAACCAAAGGGCTTGT-HCT116_Batch1 | HCT116_Batch1 | 2 | non-targeting_03016|non-targeting_03214 | Non-Targeting | 4,364 | 11,375 | 963 | 8.465934 | 1 |
[16,18,30,35,36,43,49,51,55,58,59,60,64,66,67,69,75,77,79,81,86,88,90,91,93,94,95,97,100,102,105,107(...TRUNCATED) | [1.0,1.0,12.0,5.0,10.0,1.0,2.0,10.0,3.0,1.0,3.0,7.0,4.0,27.0,6.0,17.0,1.0,3.0,6.0,6.0,7.0,3.0,1.0,2.(...TRUNCATED) | AAACCAAAGGTCCTTT-HCT116_Batch1 | HCT116_Batch1 | 2 | SBF2_P1P2-1|SBF2_P1P2-2 | SBF2 | 8,439 | 68,077 | 2,174 | 3.193443 | 1 |
[30,35,36,40,49,51,55,59,66,69,75,77,79,81,86,88,90,93,94,97,100,102,112,141,146,152,155,177,180,183(...TRUNCATED) | [2.0,2.0,7.0,2.0,1.0,9.0,2.0,1.0,4.0,10.0,3.0,1.0,1.0,5.0,2.0,2.0,1.0,1.0,3.0,2.0,1.0,1.0,3.0,1.0,2.(...TRUNCATED) | AAACCAGCAAAGCTAG-HCT116_Batch1 | HCT116_Batch1 | 2 | SLC39A8_P1P2-1|SLC39A8_P1P2-2 | SLC39A8 | 4,891 | 31,643 | 958 | 3.027526 | 1 |
[30,31,35,36,38,43,51,55,60,66,67,68,69,77,79,81,86,88,91,93,94,97,102,105,107,112,152,154,155,156,1(...TRUNCATED) | [8.0,1.0,6.0,23.0,1.0,3.0,2.0,2.0,4.0,12.0,1.0,1.0,7.0,4.0,1.0,4.0,4.0,2.0,1.0,3.0,3.0,1.0,1.0,3.0,1(...TRUNCATED) | AAACCAGCAAGCTGGG-HCT116_Batch1 | HCT116_Batch1 | 2 | CDK5RAP2_P1P2-1|CDK5RAP2_P1P2-2 | CDK5RAP2 | 6,658 | 33,785 | 883 | 2.613586 | 1 |
[18,22,30,35,36,40,42,43,51,55,60,62,64,66,67,69,75,77,79,81,86,88,90,91,93,94,105,112,124,136,146,1(...TRUNCATED) | [1.0,1.0,8.0,4.0,6.0,1.0,1.0,2.0,2.0,1.0,3.0,3.0,1.0,15.0,3.0,7.0,3.0,2.0,2.0,9.0,3.0,1.0,1.0,2.0,1.(...TRUNCATED) | AAACCAGCACTGCGTA-HCT116_Batch1 | HCT116_Batch1 | 2 | CHKA_P1P2-1|CHKA_P1P2-2 | CHKA | 7,035 | 39,414 | 1,497 | 3.798143 | 1 |
X-Atlas/Orion
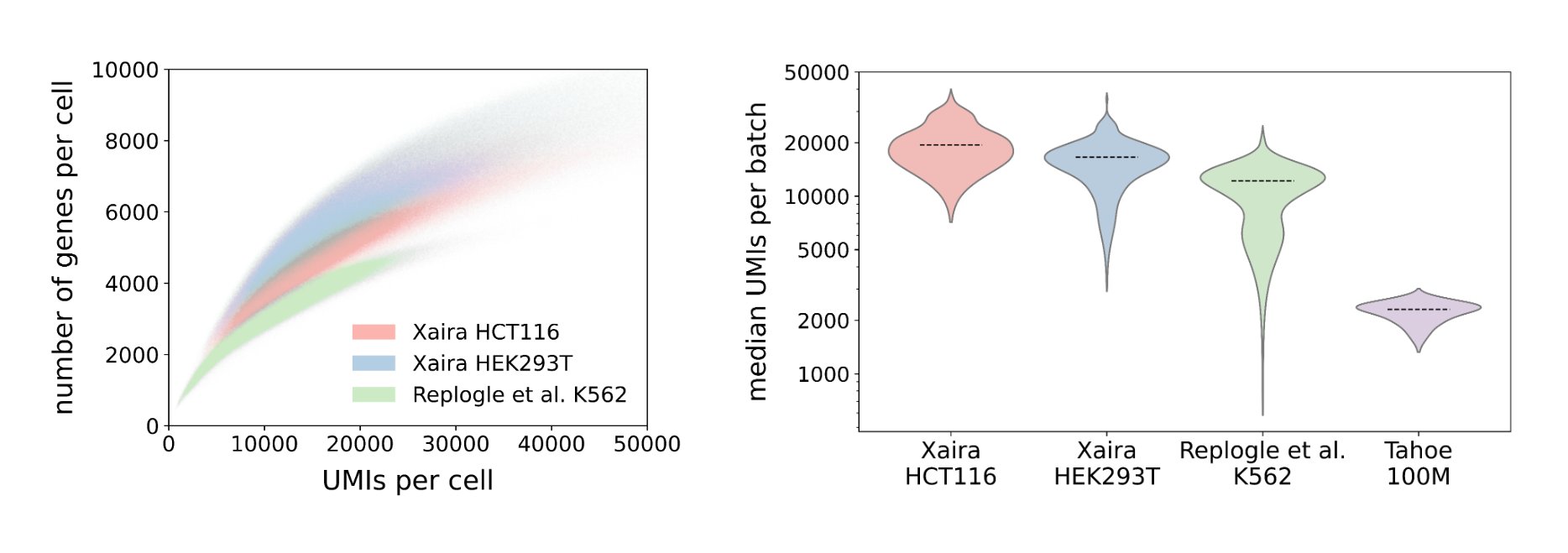
X-Atlas: Orion edition (X-Atlas/Orion) is a Perturb-seq atlas containing two genome-wide Fix-Cryopreserve-ScRNAseq (FiCS) Perturb-seq screens that target all human protein-coding genes (n = 18,903 genes). The dataset is comprised of eight million HCT116 and HEK293T cells, each deeply sequenced to a median of 16,000 unique molecular identifiers (UMIs) per cell. The median on-target knockdown efficiency is 75.4% in HCT116 cells and 51.5% in HEK293T cells, with a median of at least 140 cells per perturbation. Through the release of X-Atlas/Orion, we highlight the potential of FiCS Perturb-seq to address current scalability and variability challenges in data generation, advance foundation model development that incorporates gene-dosage effects, and accelerate biological discoveries.
Preprint: X-Atlas/Orion: Genome-wide Perturb-seq Datasets via a Scalable Fix-Cryopreserve Platform for Training Dose-Dependent Biological Foundation Models
Processed h5ads and other metadata: https://doi.org/10.25452/figshare.plus.29190726
Tutorial
from datasets import load_dataset
# load the entire dataset in streaming mode
ds = load_dataset("Xaira-Therapeutics/X-Atlas-Orion", streaming=True)
# load only hct116
hct116_ds = load_dataset("Xaira-Therapeutics/X-Atlas-Orion", streaming=True, split="HCT116")
# load only hek293t
hek293t_ds = load_dataset("Xaira-Therapeutics/X-Atlas-Orion", streaming=True, split="HEK293T")
Dataset
The dataset contains the following information:
| name | description |
|---|---|
gene_token_id |
gene identifiers corresponding to genes with non-zero expression in each cell. to be used with gene_expression. metadata/gene_metadata.parquet contains the mapping from gene_token_id to Ensembl ID and official gene symbol |
gene_expression |
raw counts for genes with non-zero expression. to be used with gene_token_id |
cell_barcode |
10X-generated cell barcode. the suffix -1 is replaced with -<SAMPLE> |
sample |
GEM batch |
num_features |
number of guides |
guide_target |
guide identity |
gene_target |
gene targeted by guide |
n_genes_by_counts |
number of genes with non-zero counts |
total_counts |
total UMIs |
total_counts_mt |
total UMIs from MT genes |
pct_counts_mt |
% UMIs from MT genes |
pass_guide_filter |
boolean if cells contains two guides from the same guide pair |
Gene metadata
All samples were aligned to the 10x Genomics GRCh38 2024-A pre-built reference genome (human reference (GRCh38) - 2024-A). Official gene symbols and ensembl IDs were extracted from the genes.gtf file.
# load metadata containing mappings to gene tokens and names
gene_metadata = load_dataset("Xaira-Therapeutics/X-Atlas-Orion","gene_metadata")
| name | description |
|---|---|
ensembl_id |
Ensembl ID |
gene_name |
official gene symbol |
gene_token_id |
gene identifiers corresponding to genes with non-zero expression in each cell. to be used with gene_token_id in the dataset |
Citation
@article{huang2025xatlasorion,
title={X-Atlas/Orion: Genome-wide Perturb-seq Datasets via a Scalable Fix-Cryopreserve Platform for Training Dose-Dependent Biological Foundation Models},
author={Huang, Ann C and Hsieh, Tsung-Han S and Zhu, Jiang and Michuda, Jackson and Teng, Ashton and Kim, Soohong and Rumsey, Elizabeth M and Lam, Sharon K and Anigbogu, Ikenna and Wright, Philip and Ameen, Mohamed and You, Kwontae and Graves, Christopher J and Kim, Hyunsung John and Litterman, Adam J and Sit, Rene V and Blocker, Alex and Chu, Ci},
journal={bioRxiv},
year={2025},
url={https://www.biorxiv.org/content/10.1101/2025.06.11.659105v1}
}
- Downloads last month
- 1,800